
As a result, priority will be given to versions of Python that include the module specified within the call to import() (i.e. versions that don’t include it will be skipped). The scanning for and binding to a version of Python typically occurs at the time of the first call to import() within an R session. For example, if you execute import("nltk") then the following locations (among other similar ones) would be scanned for a version of Python with the nltk module installed:Īt the location of the Python binary discovered on the system PATH (via the Sys.which function).Īt other customary locations for Python including /usr/local/bin/python, /opt/local/bin/python, etc. Within virtualenvs and conda envs that carry the same name as the first module imported. If specified, at the locations referenced by calls to use_python(), use_virtualenv(), and use_condaenv(). If specified, at the location referenced by the RETICULATE_PYTHON environment variable. At present, the only problem is that the drawing based on python is vague and needs to be debugged.The order in which versions of Python will be discovered and used is as follows: It is good to be familiar with a few invocation methods. Overall, in some respects, it is indeed convenient to run python directly on Rstudio. In the case of loading the reticulate package, you can directly call python. Ggplot(py$flights,aes(carrier,arr_delay)) + geom_point() Use the r object to access the object created in the R block from Python.You can use the py object to access the objects created in the Python block in R.Printable Python output, including matplotlib graphical output.At the same time shared variables/states between Python blocks. You can run Python blocks in a single Python session embedded in an R session.Reticulate includes a Python engine for R Markdown with the following functions: Of course, the shortcut keys of Rstudio are supported. Plt.bar(x + width, b, width=width, yerr = b_SD, tick_label=labels ,label='Experimental') Plt.bar(x, a, width=width, yerr = a_SD, tick_label=labels, label='Control') Use the R package, and then draw directly in Rscript: library(reticulate) In this way, you can use Pandas to read and manipulate the data, and then use ggplot2 to easily draw the Pandas data frame, although there is also a corresponding drawing method for ggplot2 in python. # Display the number of rows and columns of the data set 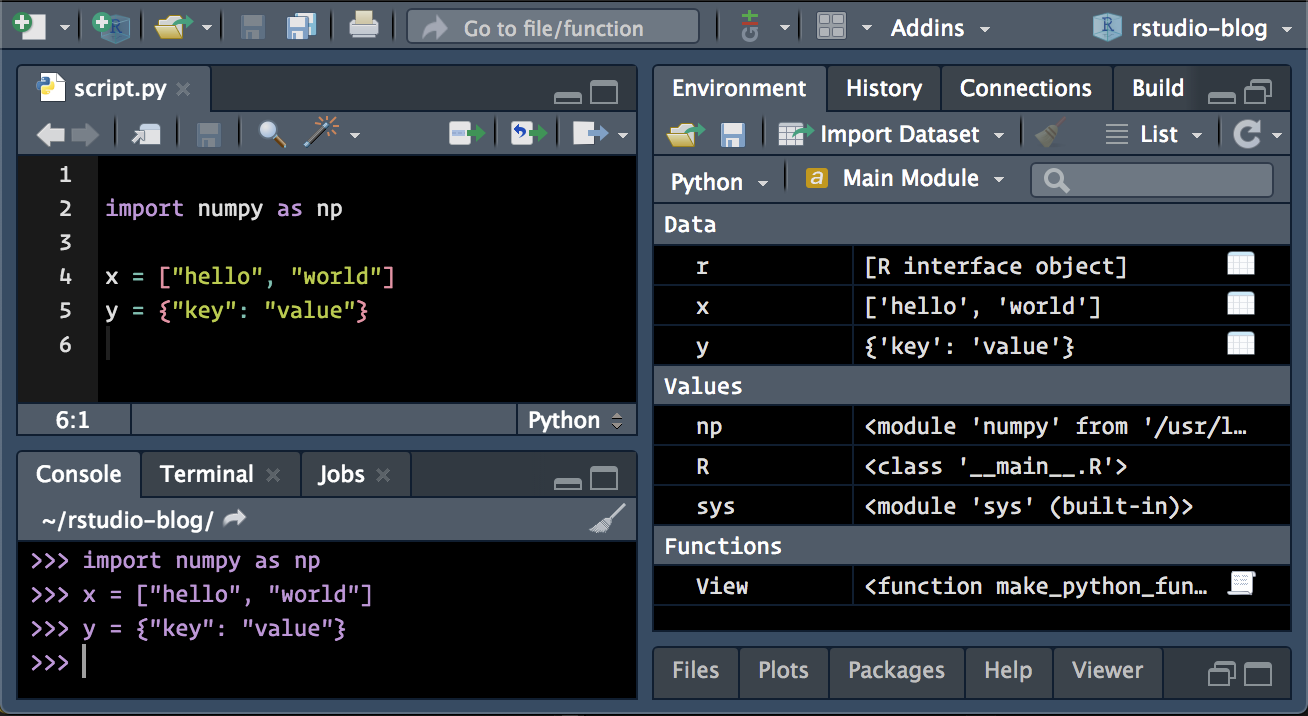
Use Python interactively ¶ #Start python command line According to different systems, there are two different operations: virtualenv is used for linux and mac and Anaconda is used for Windows. If an error occurs in the attempt, it may be caused by the conda environment. 
Py_install(packages = "numpy") #Install numpy Os$chdir("./Desktop/") #Modify working path The code is not concise enough and is not recommended. Then enter the code in the console, or use the following methods. #Specify the name of the Conda environment #Specify the directory containing Python virtualenv #Check if Python has been installed on your system, it is TRUE Test the installation environment ¶ #Load the reticulate package To run the python package in R, you must download and install it through this, which can be understood as an R-Python interface. Install the python operating environment, Anaconda is recommended.ģ. Engage your audience Create agency-quality data graphics and. If it has never been done, Rstudio will download Miniconda by default for environment construction and package management.ġ. Easily turn your data into stunning charts, maps and interactive stories. Familiar interface for R users ¶įamiliar interface, just select Python Script in New. I saw some people mentioned that the useful Python IDE is Rstudio. Pycharm has always been mentioned when we learn and test python, which is not only easy to use, but also supports the use of R language.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
